Results
- Showing results for:
- Reset all filters
Search results
-
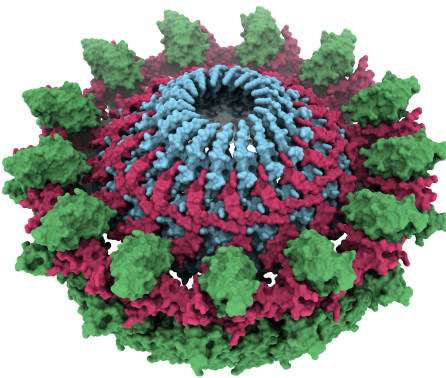
Journal articleLossi NS, Manoli E, Foerster A, et al., 2021,
The HsiB1C1 (TssB-TssC) complex of the pseudomonas aeruginosa Type VI secretion system forms a bacteriophage tail sheathlike structure
, Journal of Biological Chemistry, Vol: 288, Pages: 7536-7548, ISSN: 0021-9258Protein secretion systems in Gram-negative bacteria evolved into a variety of molecular nanomachines. They are related to cell envelope complexes, which are involved in assembly of surface appendages or transport of solutes. They are classified as types, the most recent addition being the type VI secretion system (T6SS). The T6SS displays similarities to bacteriophage tail, which drives DNA injection into bacteria. The Hcp protein is related to the T4 bacteriophage tail tube protein gp19, whereas VgrG proteins structurally resemble the gp27/gp5 puncturing device of the phage. The tube and spike of the phage are pushed through the bacterial envelope upon contraction of a tail sheath composed of gp18. In Vibrio cholerae it was proposed that VipA and VipB assemble into a tail sheathlike structure. Here we confirm these previous data by showing that HsiB1 and HsiC1 of the Pseudomonas aeruginosa H1-T6SS assemble into tubules resulting from stacking of cogwheel-like structures showing predominantly 12-fold symmetry. The internal diameter of the cogwheels is ∼100 Å, which is large enough to accommodate an Hcp tube whose external diameter has been reported to be 85 Å. The N-terminal 212 residues of HsiC1 are sufficient to form a stable complex with HsiB1, but the C terminus of HsiC1 is essential for the formation of the tubelike structure. Bioinformatics analysis suggests that HsiC1 displays similarities to gp18-like proteins in its C-terminal region. In conclusion, we provide further structural and mechanistic insights into the T6SS and show that a phage sheathlike structure is likely to be a conserved element across all T6SSs.
-
Journal articleKelwick RJR, Ricci L, Chee SM, et al., 2019,
Cell-free prototyping strategies for enhancing the sustainable production of polyhydroxyalkanoates bioplastics
, Synthetic Biology, Vol: 3, ISSN: 2397-7000The polyhydroxyalkanoates (PHAs) are microbially-produced biopolymers that could potentially be used as sustainable alternatives to oil-derived plastics. However, PHAs are currently more expensive to produce than oil-derived plastics. Therefore, more efficient production processes would be desirable. Cell-free metabolic engineering strategies have already been used to optimise several biosynthetic pathways and we envisioned that cell-free strategies could be used for optimising PHAs biosynthetic pathways. To this end, we developed several Escherichia coli cell-free systems for in vitro prototyping PHAs biosynthetic operons, and also for screening relevant metabolite recycling enzymes. Furthermore, we customised our cell-free reactions through the addition of whey permeate, an industrial waste that has been previously used to optimise in vivo PHAs production. We found that the inclusion of an optimal concentration of whey permeate enhanced relative cell-free GFPmut3b production by ∼50%. In cell-free transcription-translation prototyping reactions, GC-MS quantification of cell-free 3-hydroxybutyrate (3HB) production revealed differences between the activities of the Native ΔPhaC_C319A (1.18 ±0.39 µM), C104 ΔPhaC_C319A (4.62 ±1.31 µM) and C101 ΔPhaC_C319A (2.65 ±1.27 µM) phaCAB operons that were tested. Interestingly, the most active operon, C104 produced higher levels of PHAs (or PHAs monomers) than the Native phaCAB operon in both in vitro and in vivo assays. Coupled cell-free biotransformation/transcription-translation reactions produced greater yields of 3HB (32.87 ±6.58 µM) and these reactions were also used to characterise a Clostridium propionicum Acetyl-CoA recycling enzyme. Together, these data demonstrate that cell-free approaches complement in vivo workflows for identifying additional strategies for optimising PHAs production.
-
Conference paperGirvan P, Teng X, Brooks NJ, et al., 2019,
Redox Kinetics of the Amyloid-Beta-Copper Complex and Its Biological Implications
, 63rd Annual Meeting of the Biophysical-Society, Publisher: CELL PRESS, Pages: 28A-28A, ISSN: 0006-3495 -
Journal articleSilhan J, Zhao Q, Boura E, et al., 2018,
Structural basis for recognition and repair of the 3'-phosphate by NExo, a base excision DNA repair nuclease from Neisseria meningitidis
, Nucleic Acids Research, Vol: 46, Pages: 11980-11989, ISSN: 0305-1048NExo is an enzyme from Neisseria meningitidis that is specialized in the removal of the 3'-phosphate and other 3'-lesions, which are potential blocks for DNA repair. NExo is a highly active DNA 3'-phosphatase, and although it is from the class II AP family it lacks AP endonuclease activity. In contrast, the NExo homologue NApe, lacks 3'-phosphatase activity but is an efficient AP endonuclease. These enzymes act together to protect the meningococcus from DNA damage arising mainly from oxidative stress and spontaneous base loss. In this work, we present crystal structures of the specialized 3'-phosphatase NExo bound to DNA in the presence and absence of a 3'-phosphate lesion. We have outlined the reaction mechanism of NExo, and using point mutations we bring mechanistic insights into the specificity of the 3'-phosphatase activity of NExo. Our data provide further insight into the molecular origins of plasticity in substrate recognition for this class of enzymes. From this we hypothesize that these specialized enzymes lead to enhanced efficiency and accuracy of DNA repair and that this is important for the biological niche occupied by this bacterium.
-
Journal articleYu J, Knoppova J, Michoux F, et al., 2018,
Ycf48 involved in the biogenesis of the oxygen-evolving photosystem II complex is a seven-bladed beta-propeller protein
, Proceedings of the National Academy of Sciences, Vol: 115, Pages: E7824-E7833, ISSN: 0027-8424Robust photosynthesis in chloroplasts and cyanobacteria requires the participation of accessory proteins to facilitate the assembly and maintenance of the photosynthetic apparatus located within the thylakoid membranes. The highly conserved Ycf48 protein acts early in the biogenesis of the oxygen-evolving photosystem II (PSII) complex by binding to newly synthesized precursor D1 subunit and by promoting efficient association with the D2 protein to form a PSII reaction center (PSII RC) assembly intermediate. Ycf48 is also required for efficient replacement of damaged D1 during the repair of PSII. However, the structural features underpinning Ycf48 function remain unclear. Here we show that Ycf48 proteins encoded by the thermophilic cyanobacterium Thermosynechococcus elongatus and the red alga Cyanidioschyzon merolae form seven-bladed beta-propellers with the 19-aa insertion characteristic of eukaryotic Ycf48 located at the junction of blades 3 and 4. Knowledge of these structures has allowed us to identify a conserved “Arg patch” on the surface of Ycf48 that is important for binding of Ycf48 to PSII RCs but also to larger complexes, including trimeric photosystem I (PSI). Reduced accumulation of chlorophyll in the absence of Ycf48 and the association of Ycf48 with PSI provide evidence of a more wide-ranging role for Ycf48 in the biogenesis of the photosynthetic apparatus than previously thought. Copurification of Ycf48 with the cyanobacterial YidC protein insertase supports the involvement of Ycf48 during the cotranslational insertion of chlorophyll-binding apopolypeptides into the membrane.
-
Journal articleBernal P, Llamas MA, Filloux A, 2018,
Type VI secretion systems in plant-associated bacteria
, Environmental Microbiology, Vol: 20, Pages: 1-15, ISSN: 1462-2912The type VI secretion system (T6SS) is a bacterial nanomachine used to inject effectors into prokaryotic or eukaryotic cells and is thus involved in both host manipulation and interbacterial competition. The T6SS is widespread among Gram‐negative bacteria, mostly within the Proteobacterium Phylum. This secretion system is commonly found in commensal and pathogenic plant‐associated bacteria. Phylogenetic analysis of phytobacterial T6SS clusters shows that they are distributed in the five main clades previously described (group 1–5). The even distribution of the system among commensal and pathogenic phytobacteria suggests that the T6SS provides fitness and colonization advantages in planta and that the role of the T6SS is not restricted to virulence. This manuscript reviews the phylogeny and biological roles of the T6SS in plant‐associated bacteria, highlighting a remarkable diversity both in terms of mechanism and function.
-
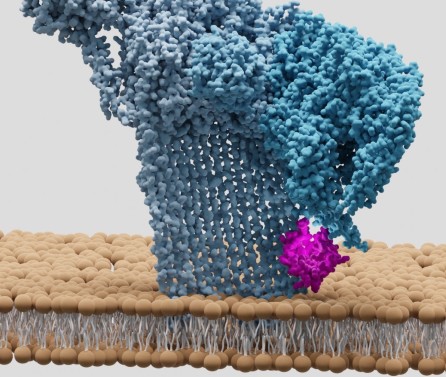
Journal articleFreemont PS, Salih O, He S, et al., 2018,
Atomic Structure of Type VI Contractile Sheath from Pseudomonas aeruginosa
, Structure, Vol: 26, Pages: 329-336.e3, ISSN: 0969-2126Pseudomonas aeruginosa has three type VI secretion systems (T6SSs), H1-, H2-, and H3-T6SS, each belonging to a distinct group. The two T6SS components, TssB/VipA and TssC/VipB, assemble to form tubules that conserve structural/functional homology with tail sheaths of contractile bacteriophages and pyocins. Here, we used cryoelectron microscopy to solve the structure of the H1-T6SS P. aeruginosa TssB1C1 sheath at 3.3 Å resolution. Our structure allowed us to resolve some features of the T6SS sheath that were not resolved in the Vibrio cholerae VipAB and Francisella tularensis IglAB structures. Comparison with sheath structures from other contractile machines, including T4 phage and R-type pyocins, provides a better understanding of how these systems have conserved similar functions/mechanisms despite evolution. We used the P. aeruginosa R2 pyocin as a structural template to build an atomic model of the TssB1C1 sheath in its extended conformation, allowing us to propose a coiled-spring-like mechanism for T6SS sheath contraction.
-
Journal articleMullineaux-Sanders C, Colins JW, Ruano-Gallego D, et al., 2017,
Citrobacter rodentium relies on commensals for colonization of the colonic mucosa
, Cell Reports, Vol: 21, Pages: 3381-3389, ISSN: 2211-1247We investigated the role of commensals at the peak of infection with the colonic mouse pathogen Citrobacter rodentium. Bioluminescent and kanamycin (Kan)-resistant C. rodentium persisted avirulently in the cecal lumen of mice continuously treated with Kan. A single Kan treatment was sufficient to displace C. rodentium from the colonic mucosa, a phenomenon not observed following treatment with vancomycin (Van) or metronidazole (Met). Kan, Van, and Met induce distinct dysbiosis, suggesting C. rodentium relies on specific commensals for colonic colonization. Expression of the master virulence regulator ler is induced in germ-free mice, yet C. rodentium is only seen in the cecal lumen. Moreover, in conventional mice, a single Kan treatment was sufficient to displace C. rodentium constitutively expressing Ler from the colonic mucosa. These results show that expression of virulence genes is not sufficient for colonization of the colonic mucosa and that commensals are essential for a physiological infection course.
-
Journal articleJohnson R, Ravenhall M, Pickard D, et al., 2017,
Comparison of Salmonella enterica serovars Typhi and Typhimurium reveals typhoidal-specific responses to bile
, Infection and Immunity, Vol: 86, ISSN: 0019-9567Salmonella enterica serovars Typhi and Typhimurium cause typhoid fever and gastroenteritis respectively. A unique feature of typhoid infection is asymptomatic carriage within the gallbladder, which is linked with S. Typhi transmission. Despite this, S. Typhi responses to bile have been poorly studied. RNA-Seq of S. Typhi Ty2 and a clinical S. Typhi isolate belonging to the globally dominant H58 lineage (129-0238), as well as S. Typhimurium 14028, revealed that 249, 389 and 453 genes respectively were differentially expressed in the presence of 3% bile compared to control cultures lacking bile. fad genes, the actP-acs operon, and putative sialic acid uptake and metabolism genes (t1787-t1790) were upregulated in all strains following bile exposure, which may represent adaptation to the small intestine environment. Genes within the Salmonella pathogenicity island 1 (SPI-1), encoding a type IIII secretion system (T3SS), and motility genes were significantly upregulated in both S. Typhi strains in bile, but downregulated in S. Typhimurium. Western blots of the SPI-1 proteins SipC, SipD, SopB and SopE validated the gene expression data. Consistent with this, bile significantly increased S. Typhi HeLa cell invasion whilst S. Typhimurium invasion was significantly repressed. Protein stability assays demonstrated that in S. Typhi the half-life of HilD, the dominant regulator of SPI-1, is three times longer in the presence of bile; this increase in stability was independent of the acetyltransferase Pat. Overall, we found that S. Typhi exhibits a specific response to bile, especially with regards to virulence gene expression, which could impact pathogenesis and transmission.
-
Journal articleZhang X, Aramayo RJ, Willhoft O, et al., 2017,
CryoEM structures of the human INO80 chromatin remodelling complex
, Nature Structural and Molecular Biology, Vol: 25, Pages: 37-44, ISSN: 1545-9985Access to chromatin for processes such as DNA repair and transcription requires the sliding of nucleosomes along DNA. The multi-subunit INO80 chromatin remodelling complex has a particular role in DNA repair. Here we present the cryo electron microscopy structures of the active core complex of human INO80 at 9.6 Å with portions at 4.1 Å resolution along with reconstructions of combinations of subunits. Together these structures reveal the architecture of the INO80 complex, including Ino80 and actin-related proteins, which is assembled around a single Tip49a (RUVBL1) and Tip49b (RUVBL2) AAA+ heterohexamer. An unusual spoked-wheel structural domain of the Ino80 subunit is engulfed by this heterohexamer and the intimate association of this Ino80 domain with the heterohexamer is at the core of the complex. We also identify a cleft in RUVBL1 and RUVBL2, which forms a major interaction site for partner proteins and likely communicates partner-interactions with its nucleotide binding sites.
-
Journal articlePortaliou AG, Tsolis KC, Loos MS, et al., 2017,
Hierarchical protein targeting and secretion is controlled by an affinity switch in the type III secretion system of enteropathogenic <i>Escherichia coli</i>
, EMBO JOURNAL, Vol: 36, Pages: 3517-3531, ISSN: 0261-4189- Author Web Link
- Cite
- Citations: 43
-
Journal articleCepeda-Molero M, Berger CN, Walsham ADS, et al., 2017,
Attaching and effacing (A/E) lesion formation by enteropathogenic E. coli on human intestinal mucosa is dependent on non-LEE effectors
, PLoS Pathogens, Vol: 13, ISSN: 1553-7366Enteropathogenic E. coli (EPEC) is a human pathogen that causes acute and chronic pediatric diarrhea. The hallmark of EPEC infection is the formation of attaching and effacing (A/E) lesions in the intestinal epithelium. Formation of A/E lesions is mediated by genes located on the pathogenicity island locus of enterocyte effacement (LEE), which encode the adhesin intimin, a type III secretion system (T3SS) and six effectors, including the essential translocated intimin receptor (Tir). Seventeen additional effectors are encoded by genes located outside the LEE, in insertion elements and prophages. Here, using a stepwise approach, we generated an EPEC mutant lacking the entire effector genes (EPEC0) and intermediate mutants. We show that EPEC0 contains a functional T3SS. An EPEC mutant expressing intimin but lacking all the LEE effectors but Tir (EPEC1) was able to trigger robust actin polymerization in HeLa cells and mucin-producing intestinal LS174T cells. However, EPEC1 was unable to form A/E lesions on human intestinal in vitro organ cultures (IVOC). Screening the intermediate mutants for genes involved in A/E lesion formation on IVOC revealed that strains lacking non-LEE effector/s have a marginal ability to form A/E lesions. Furthermore, we found that Efa1/LifA proteins are important for A/E lesion formation efficiency in EPEC strains lacking multiple effectors. Taken together, these results demonstrate the intricate relationships between T3SS effectors and the essential role non-LEE effectors play in A/E lesion formation on mucosal surfaces.
-
Journal articleNoguchi Y, yuan Z, Bai L, et al., 2017,
Cryo-EM structure of Mcm2-7 double-hexamer on DNA suggests a lagging strand DNA extrusion model
, Proceedings of the National Academy of Sciences, Vol: 114, Pages: E9529-E9538, ISSN: 0027-8424During replication initiation, the core component of the helicase—the Mcm2-7 hexamer—is loaded on origin DNA as a double hexamer (DH). The two ring-shaped hexamers are staggered, leading to a kinked axial channel. How the origin DNA interacts with the axial channel is not understood, but the interaction could provide key insights into Mcm2-7 function and regulation. Here, we report the cryo-EM structure of the Mcm2-7 DH on dsDNA and show that the DNA is zigzagged inside the central channel. Several of the Mcm subunit DNA-binding loops, such as the oligosaccharide–oligonucleotide loops, helix 2 insertion loops, and presensor 1 (PS1) loops, are well defined, and many of them interact extensively with the DNA. The PS1 loops of Mcm 3, 4, 6, and 7, but not 2 and 5, engage the lagging strand with an approximate step size of one base per subunit. Staggered coupling of the two opposing hexamers positions the DNA right in front of the two Mcm2–Mcm5 gates, with each strand being pressed against one gate. The architecture suggests that lagging-strand extrusion initiates in the middle of the DH that is composed of the zinc finger domains of both hexamers. To convert the Mcm2-7 DH structure into the Mcm2-7 hexamer structure found in the active helicase, the N-tier ring of the Mcm2-7 hexamer in the DH-dsDNA needs to tilt and shift laterally. We suggest that these N-tier ring movements cause the DNA strand separation and lagging-strand extrusion.
-
Journal articleWen KY, Cameron L, Chappell J, et al., 2017,
A Cell-Free Biosensor for Detecting Quorum Sensing Molecules in P. aeruginosa-Infected Respiratory Samples.
, ACS Synthetic Biology, Vol: 6, Pages: 2293-2301, ISSN: 2161-5063Synthetic biology designed cell-free biosensors are a promising new tool for the detection of clinically relevant biomarkers in infectious diseases. Here, we report that a modular DNA-encoded biosensor in cell-free protein expression systems can be used to measure a bacterial biomarker of Pseudomonas aeruginosa infection from human sputum samples. By optimizing the cell-free system and sample extraction, we demonstrate that the quorum sensing molecule 3-oxo-C12-HSL in sputum samples from cystic fibrosis lungs can be quantitatively measured at nanomolar levels using our cell-free biosensor system, and is comparable to LC-MS measurements of the same samples. This study further illustrates the potential of modular cell-free biosensors as rapid, low-cost detection assays that can inform clinical practice.
-
Journal articleBerger C, Crepin V, Roumeliotis TI, et al., 2017,
Citrobacter rodentium subverts ATP flux 1 and cholesterol homeostasis in 2 intestinal epithelial cell in vivo
, Cell Metabolism, Vol: 26, Pages: 738-752.e6, ISSN: 1550-4131The intestinal epithelial cells (IECs) that line the gut form a robust line of defense against ingested pathogens. We investigated the impact of infection with the enteric pathogen Citrobacter rodentium on mouse IEC metabolism using global proteomic and targeted metabolomics and lipidomics. The major signatures of the infection were upregulation of the sugar transporter Sglt4, aerobic glycolysis, and production of phosphocreatine, which mobilizes cytosolic energy. In contrast, biogenesis of mitochondrial cardiolipins, essential for ATP production, was inhibited, which coincided with increased levels of mucosal O2 and a reduction in colon-associated anaerobic commensals. In addition, IECs responded to infection by activating Srebp2 and the cholesterol biosynthetic pathway. Unexpectedly, infected IECs also upregulated the cholesterol efflux proteins AbcA1, AbcG8, and ApoA1, resulting in higher levels of fecal cholesterol and a bloom of Proteobacteria. These results suggest that C. rodentium manipulates host metabolism to evade innate immune responses and establish a favorable gut ecosystem.
-
Journal articleMcCarthy RR, Valentini M, Filloux A, 2017,
Contribution of Cyclic di-GMP in the Control of Type III and Type VI Secretion in Pseudomonas aeruginosa.
, Methods Mol Biol, Vol: 1657, Pages: 213-224Bacteria produce toxins to enhance their competitiveness in the colonization of an environment as well as during an infection. The delivery of toxins into target cells is mediated by several types of secretion systems, among them our focus is Type III and Type VI Secretion Systems (T3SS and T6SS, respectively). A thorough methodology is provided detailing how to identify if cyclic di-GMP signaling plays a role in the P. aeruginosa toxin delivery mediated by T3SS or T6SS. This includes in vitro preparation of the samples for Western blot analysis aiming at detecting possible c-di-GMP-dependent T3SS/T6SS switch, as well as in vivo analysis using the model organism Galleria mellonella to demonstrate the ecological and pathogenic consequence of the switch between these two secretion systems.
-
Journal articleKarampatzakis A, Song CZ, Allsopp LP, et al., 2017,
Probing the internal micromechanical properties of Pseudomonas aeruginosa biofilms by Brillouin imaging.
, NPJ Biofilms Microbiomes, Vol: 3, ISSN: 2055-5008Biofilms are organised aggregates of bacteria that adhere to each other or surfaces. The matrix of extracellular polymeric substances that holds the cells together provides the mechanical stability of the biofilm. In this study, we have applied Brillouin microscopy, a technique that is capable of measuring mechanical properties of specimens on a micrometre scale based on the shift in frequency of light incident upon a sample due to thermal fluctuations, to investigate the micromechanical properties of an active, live Pseudomonas aeruginosa biofilm. Using this non-contact and label-free technique, we have extracted information about the internal stiffness of biofilms under continuous flow. No correlation with colony size was found when comparing the averages of Brillouin shifts of two-dimensional cross-sections of randomly selected colonies. However, when focusing on single colonies, we observed two distinct spatial patterns: in smaller colonies, stiffness increased towards their interior, indicating a more compact structure of the centre of the colony, whereas, larger (over 45 μm) colonies were found to have less stiff interiors.
-
Journal articleHeeb S, Camara M, Filloux A, et al., 2017,
Professor Dieter Haas (1945-2017)
, FEMS Microbiology Reviews, Vol: 41, Pages: 597-598, ISSN: 0168-6445 -
Journal articleJakobsen TH, Warming AN, Vejborg RM, et al., 2017,
A broad range quorum sensing inhibitor working through sRNA inhibition
, Scientific Reports, Vol: 7, ISSN: 2045-2322For the last decade, chemical control of bacterial virulence has received considerable attention. Ajoene, a sulfur-rich molecule from garlic has been shown to reduce expression of key quorum sensing regulated virulence factors in the opportunistic pathogen Pseudomonas aeruginosa. Here we show that the repressing effect of ajoene on quorum sensing occurs by inhibition of small regulatory RNAs (sRNA) in P. aeruginosa as well as in Staphylococcus aureus, another important human pathogen that employs quorum sensing to control virulence gene expression. Using various reporter constructs, we found that ajoene lowered expression of the sRNAs RsmY and RsmZ in P. aeruginosa and the small dual-function regulatory RNA, RNAIII in S. aureus, that controls expression of key virulence factors. We confirmed the modulation of RNAIII by RNA sequencing and found that the expression of many QS regulated genes encoding virulence factors such as hemolysins and proteases were lowered in the presence of ajoene in S. aureus. Importantly, our findings show that sRNAs across bacterial species potentially may qualify as targets of anti-virulence therapy and that ajoene could be a lead structure in search of broad-spectrum compounds transcending the Gram negative-positive borderline.
-
Journal articlefreemont P, Stach L, 2017,
The AAA+ ATPase p97, a cellular multi-tool
, Biochemical Journal, Vol: 474, Pages: 2953-2976, ISSN: 1470-8728The AAA+ (ATPases associated with diverse cellular activities) ATPase p97 is essential to a wide range of cellular functions, including endoplasmic reticulum-associated degradation, membrane fusion, NF-κB (nuclear factor kappa-light-chain-enhancer of activated B cells) activation and chromatin-associated processes, which are regulated by ubiquitination. p97 acts downstream from ubiquitin signaling events and utilizes the energy from ATP hydrolysis to extract its substrate proteins from cellular structures or multiprotein complexes. A multitude of p97 cofactors have evolved which are essential to p97 function. Ubiquitin-interacting domains and p97-binding domains combine to form bi-functional cofactors, whose complexes with p97 enable the enzyme to interact with a wide range of ubiquitinated substrates. A set of mutations in p97 have been shown to cause the multisystem proteinopathy inclusion body myopathy associated with Paget's disease of bone and frontotemporal dementia. In addition, p97 inhibition has been identified as a promising approach to provoke proteotoxic stress in tumors. In this review, we will describe the cellular processes governed by p97, how the cofactors interact with both p97 and its ubiquitinated substrates, p97 enzymology and the current status in developing p97 inhibitors for cancer therapy.
-
Journal articleBeckova M, Yu J, Krynicka V, et al., 2017,
Structure of Psb29/Thf1 and its association with the FtsH protease complex involved in photosystem II repair in cyanobacteria
, Philosophical Transactions of the Royal Society B: Biological Sciences, Vol: 372, ISSN: 1471-2970One strategy for enhancing photosynthesis in crop plants is to improve the ability to repair photosystem II (PSII) in response to irreversible damage by light. Despite the pivotal role of thylakoid embedded FtsH protease complexes in the selective degradation of PSII subunits during repair, little is known about the factors involved in regulating FtsH expression. Here we show using the cyanobacterium Synechocystis sp. PCC 6803 that the Psb29 subunit, originally identified as a minor component of His tagged PSII preparations, physically interacts with FtsH complexes in vivo and is required for normal accumulation of the FtsH2/FtsH3 hetero oligomeric complex involved in PSII repair. We show using X ray crystallography that Psb29 from Thermosynechococcus elongatushas a unique fold consisting of a helical bundle and an extended C terminal helix and contains a highly conserved region that might be involved in binding to FtsH. A similar interaction is likely to occur in Arabidopsis chloroplasts between the Psb29 homologue, termed THF1, and the FTSH2/FTSH5 complex. The direct involvement of Psb29/THF1 in FtsH accumulation helps explain why THF1 is a target during the hypersensitive response in plants induced by pathogen infection. Downregulating FtsH function and the PSII repair cycle via THF1 would contribute to the production
-
Journal articleDavies SK, Fearn S, Allsopp LP, et al., 2017,
Visualizing Antimicrobials in BacterialBiofilms: Three-Dimensional BiochemicalImaging Using TOF-SIMS
, mSphere, Vol: 2, ISSN: 2379-5042Bacterial biofilms are groups of bacteria that exist within a self-produced extracellular matrix, adhering to each other and usually to a surface. They grow on medical equipment and inserts such as catheters and are responsible for many persistent infections throughout the body, as they can have high resistance to many antimicrobials. Pseudomonas aeruginosa is an opportunistic pathogen that can cause both acute and chronic infections and is used as a model for research into biofilms. Direct biochemical methods of imaging of molecules in bacterial biofilms are of high value in gaining a better understanding of the fundamental biology of biofilms and biochemical gradients within them. Time of flight–secondary-ion mass spectrometry (TOF-SIMS) is one approach, which combines relatively high spatial resolution and sensitivity and can perform depth profiling analysis. It has been used to analyze bacterial biofilms but has not yet been used to study the distribution of antimicrobials (including antibiotics and the antimicrobial metal gallium) within biofilms. Here we compared two methods of imaging of the interior structure of P. aeruginosa in biological samples using TOF-SIMS, looking at both antimicrobials and endogenous biochemicals: cryosectioning of tissue samples and depth profiling to give pseudo-three-dimensional (pseudo-3D) images. The sample types included both simple biofilms grown on glass slides and bacteria growing in tissues in an ex vivo pig lung model. The two techniques for the 3D imaging of biofilms are potentially valuable complementary tools for analyzing bacterial infection.
-
Journal articleAllsopp LP, Wood TE, Howard SA, et al., 2017,
RsmA and AmrZ orchestrate the assembly of all three type VI secretion systems in Pseudomonas aeruginosa
, Proceedings of the National Academy of Sciences of the United States of America, Vol: 114, Pages: 7707-7712, ISSN: 1091-6490The type VI secretion system (T6SS) is a weapon of bacterial warfare and host cell subversion. The Gram-negative pathogen Pseudomonas aeruginosa has three T6SSs involved in colonization, competition, and full virulence. H1-T6SS is a molecular gun firing seven toxins, Tse1–Tse7, challenging survival of other bacteria and helping P. aeruginosa to prevail in specific niches. The H1-T6SS characterization was facilitated through studying a P. aeruginosa strain lacking the RetS sensor, which has a fully active H1-T6SS, in contrast to the parent. However, study of H2-T6SS and H3-T6SS has been neglected because of a poor understanding of the associated regulatory network. Here we performed a screen to identify H2-T6SS and H3-T6SS regulatory elements and found that the posttranscriptional regulator RsmA imposes a concerted repression on all three T6SS clusters. A higher level of complexity could be observed as we identified a transcriptional regulator, AmrZ, which acts as a negative regulator of H2-T6SS. Overall, although the level of T6SS transcripts is fine-tuned by AmrZ, all T6SS mRNAs are silenced by RsmA. We expanded this concept of global control by RsmA to VgrG spike and T6SS toxin transcripts whose genes are scattered on the chromosome. These observations triggered the characterization of a suite of H2-T6SS toxins and their implication in direct bacterial competition. Our study thus unveils a central mechanism that modulates the deployment of all T6SS weapons that may be simultaneously produced within a single cell.
-
Journal articleGlyde R, Ye F, Darbari V, et al., 2017,
Structures of RNA polymerase closed and intermediate complexes revealmechanisms of DNA opening and transcription initiation
, Molecular Cell, Vol: 67, Pages: 106-116, ISSN: 1097-2765Gene transcription is carried out by RNA polymerases (RNAPs). For transcription to occur, the closed promoter complex (RPc), where DNA is double stranded, must isomerize into an open promoter complex (RPo), where the DNA is melted out into a transcription bubble and the single-stranded template DNA is delivered to the RNAP active site. Using a bacterial RNAP containing the alternative σ54 factor and cryoelectron microscopy, we determined structures of RPc and the activator-bound intermediate complex en route to RPo at 3.8 and 5.8 Å. Our structures show how RNAP-σ54 interacts with promoter DNA to initiate the DNA distortions required for transcription bubble formation, and how the activator interacts with RPc, leading to significant conformational changes in RNAP and σ54 that promote RPo formation. We propose that DNA melting is an active process initiated in RPc and that the RNAP conformations of intermediates are significantly different from that of RPc and RPo.
-
Journal articleBerger CN, 2017,
The Enterohemorrhagic Escherichia coli Effector EspW Triggers Actin Remodeling in a Rac1-Dependent Manner
, Infection and Immunity, Vol: 85, ISSN: 1098-5522Enterohemorrhagic Escherichia coli (EHEC) is a diarrheagenic pathogen that colonizes the gut mucosa and induces attaching-and-effacing lesions. EHEC employs a type III secretion system (T3SS) to translocate 50 effector proteins that hijack and manipulate host cell signaling pathways, which allow bacterial colonization and subversion of immune responses and disease progression. The aim of this study was to characterize the T3SS effector EspW. We found espW in the sequenced O157:H7 and non-O157 EHEC strains as well as in Shigella boydii. Furthermore, a truncated version of EspW, containing the first 206 residues, is present in EPEC strains belonging to serotype O55:H7. Screening a collection of clinical EPEC isolates revealed that espW is present in 52% of the tested strains. We report that EspW modulates actin dynamics in a Rac1-dependent manner. Ectopic expression of EspW results in formation of unique membrane protrusions. Infection of Swiss cells with an EHEC espW deletion mutant induces a cell shrinkage phenotype that could be rescued by Rac1 activation via expression of the bacterial guanine nucleotide exchange factor, EspT. Furthermore, using a yeast two-hybrid screen, we identified the motor protein Kif15 as a potential interacting partner of EspW. Kif15 and EspW colocalized in cotransfected cells, while ectopically expressed Kif15 localized to the actin pedestals following EHEC infection. The data suggest that Kif15 recruits EspW to the site of bacterial attachment, which in turn activates Rac1, resulting in modifications of the actin cytoskeleton that are essential to maintain cell shape during infection.
-
Journal articleSmith WD, Bardin E, Cameron L, et al., 2017,
Current and future therapies for Pseudomonas aeruginosa infection in patients with cystic fibrosis
, FEMS Microbiology Letters, Vol: 364, ISSN: 0378-1097Pseudomonas aeruginosa opportunistically infects the airways of patients with cystic fibrosis and causes significant morbidity and mortality. Initial infection can often be eradicated though requires prompt detection and adequate treatment. Intermittent and then chronic infection occurs in the majority of patients. Better detection of P. aeruginosa infection using biomarkers may enable more successful eradication before chronic infection is established. In chronic infection P. aeruginosa adapts to avoid immune clearance and resist antibiotics via efflux pumps, β-lactamase expression, reduced porins and switching to a biofilm lifestyle. The optimal treatment strategies for P. aeruginosa infection are still being established, and new antibiotic formulations such as liposomal amikacin, fosfomycin in combination with tobramycin and inhaled levofloxacin are being explored. Novel agents such as the alginate oligosaccharide OligoG, cysteamine, bacteriophage, nitric oxide, garlic oil and gallium may be useful as anti-pseudomonal strategies, and immunotherapy to prevent infection may have a role in the future. New treatments that target the primary defect in cystic fibrosis, recently licensed for use, have been associated with a fall in P. aeruginosa infection prevalence. Understanding the mechanisms for this could add further strategies for treating P. aeruginosa in future.
-
Journal articleWigley DB, Willhoft O, McCormack EA, et al., 2017,
Cross-talk within a functional INO80 complex dimer regulates nucleosome sliding
, eLife, Vol: 6, ISSN: 2050-084XSeveral chromatin remodellers have the ability to space nucleosomes on DNA. For ISWI remodellers, this involves an interplay between H4 histone tails, the AutoN and NegC motifs of the motor domains that together regulate ATPase activity and sense the length of DNA flanking the nucleosome. By contrast, the INO80 complex also spaces nucleosomes but is not regulated by H4 tails and lacks the AutoN and NegC motifs. Instead nucleosome sliding requires cooperativity between two INO80 complexes that monitor DNA length simultaneously on either side of the nucleosome during sliding. The C-terminal domain of the human Ino80 subunit (Ino80CTD) binds cooperatively to DNA and dimerisation of these domains provides crosstalk between complexes. ATPase activity, rather than being regulated, instead gradually becomes uncoupled as nucleosome sliding reaches an end point and this is controlled by the Ino80CTD. A single active ATPase motor within the dimer is sufficient for sliding.
-
Journal articleWitcomb LA, Czupryna J, Francis KP, et al., 2017,
Non-invasive three-dimensional imaging of Escherichia coli K1 infection using diffuse light imaging tomography combined with micro-computed tomography
, Methods, Vol: 127, Pages: 62-68, ISSN: 1046-2023In contrast to two-dimensional bioluminescence imaging, three dimensional diffuse light imaging tomography with integrated micro-computed tomography (DLIT-μCT) has the potential to realise spatial variations in infection patterns when imaging experimental animals dosed with derivatives of virulent bacteria carrying bioluminescent reporter genes such as the lux operon from the bacterium Photorhabdus luminescens. The method provides an opportunity to precisely localise the bacterial infection sites within the animal and enables the generation of four-dimensional movies of the infection cycle. Here, we describe the use of the PerkinElmer IVIS SpectrumCT in vivo imaging system to investigate progression of lethal systemic infection in neonatal rats following colonisation of the gastrointestinal tract with the neonatal pathogen Escherichia coli K1. We confirm previous observations that these bacteria stably colonize the colon and small intestine following feeding of the infectious dose from a micropipette; invading bacteria migrate across the gut epithelium into the blood circulation and establish foci of infection in major organs, including the brain. DLIT-μCT revealed novel multiple sites of colonisation within the alimentary canal, including the tongue, oesophagus and stomach, with penetration of the non-keratinised oesophageal epithelial surface, providing strong evidence of a further major site for bacterial dissemination. We highlight technical issues associated with imaging of infections in new born rat pups and show that the whole-body and organ bioburden correlates with disease severity.
-
Journal articleImbert PRC, Louche A, Luizet J-B, et al., 2017,
A Pseudomonas aeruginosa TIR effector mediates immune evasion by targeting UBAP1 and TLR adaptors
, EMBO JOURNAL, Vol: 36, Pages: 1869-1887, ISSN: 0261-4189 -
Journal articlePallett MA, Crepin VF, Serafini N, et al., 2017,
Bacterial virulence factor inhibits caspase-4/11 activation in intestinal epithelial cells
, Mucosal Immunology, Vol: 10, Pages: 602-612, ISSN: 1935-3456The human pathogen enteropathogenic Escherichia coli (EPEC), as well as the mouse pathogen Citrobacter rodentium, colonize the gut mucosa via attaching and effacing lesion formation and cause diarrheal diseases. EPEC and C. rodentium type III secretion system (T3SS) effectors repress innate immune responses and infiltration of immune cells. Inflammatory caspases such as caspase-1 and caspase-4/11 are crucial mediators of host defense and inflammation in the gut via their ability to process cytokines such as interleukin (IL)-1β and IL-18. Here we report that the effector NleF binds the catalytic domain of caspase-4 and inhibits its proteolytic activity. Following infection of intestinal epithelial cells (IECs) EPEC inhibited caspase-4 and IL-18 processing in an NleF-dependent manner. Depletion of caspase-4 in IECs prevented the secretion of mature IL-18 in response to infection with EPECΔnleF. NleF-dependent inhibition of caspase-11 in colons of mice prevented IL-18 secretion and neutrophil influx at early stages of C. rodentium infection. Neither wild-type C. rodentium nor C. rodentiumΔnleF triggered neutrophil infiltration or IL-18 secretion in Cas11 or Casp1/11-deficient mice. Thus, IECs have a key role in modulating early innate immune responses in the gut via a caspase-4/11—IL-18 axis, which is targeted by virulence factors encoded by enteric pathogens.
This data is extracted from the Web of Science and reproduced under a licence from Thomson Reuters. You may not copy or re-distribute this data in whole or in part without the written consent of the Science business of Thomson Reuters.
Centre for Structural Biology Open Day
Join us for our Open Day on 16 May 2024 - find out more!